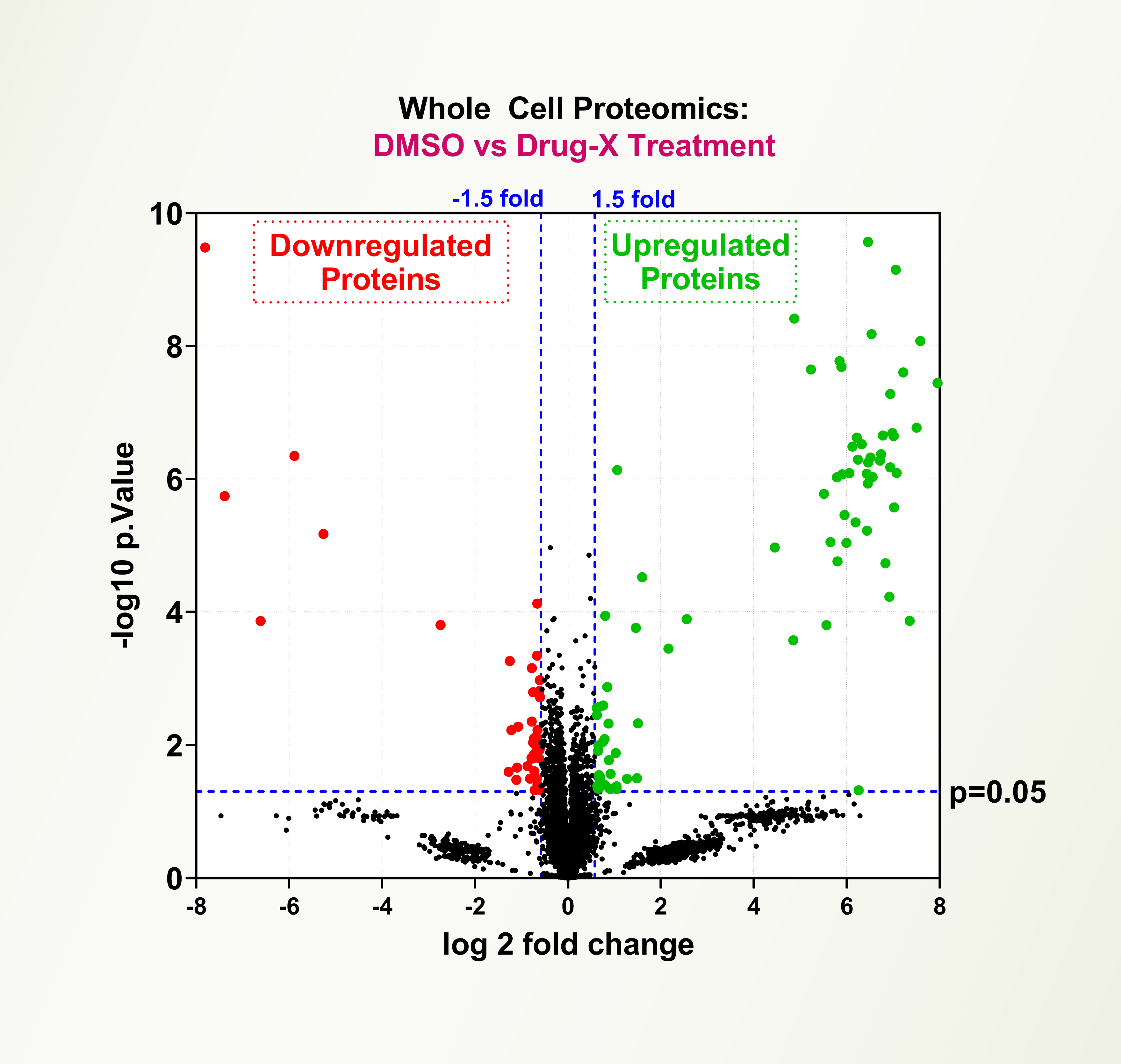
Traditional methods for mass spectrometry-based proteomics, such as SILAC (stable isotope labeling with amino acids in cell culture), are insensitive, poorly reproducible, costly, and time consuming. LifeSensors’ TUBEs bind to all polyubiquitin chains with 1-10 nM affinity, overcoming major problems common to mass spectrometry proteomics. LifeSensors has developed K63-, K48-, and M1(linear)-chain selective TUBEs. LifeSensors is actively engaged in developing K6, K11, K29, K27, and K33 polyubiquitin chain-selective TUBEs. The combination of the TUBE-based affinity technology and targeted mass spectrometry is the most powerful way to detect the alternations in PMTs and identify signatures for research and biomarkers. LifeSensors’ ubiquitin proteomics technology can detect ultra-low levels of ubiquitylated biomarkers from tissues and cells.
Associated Services
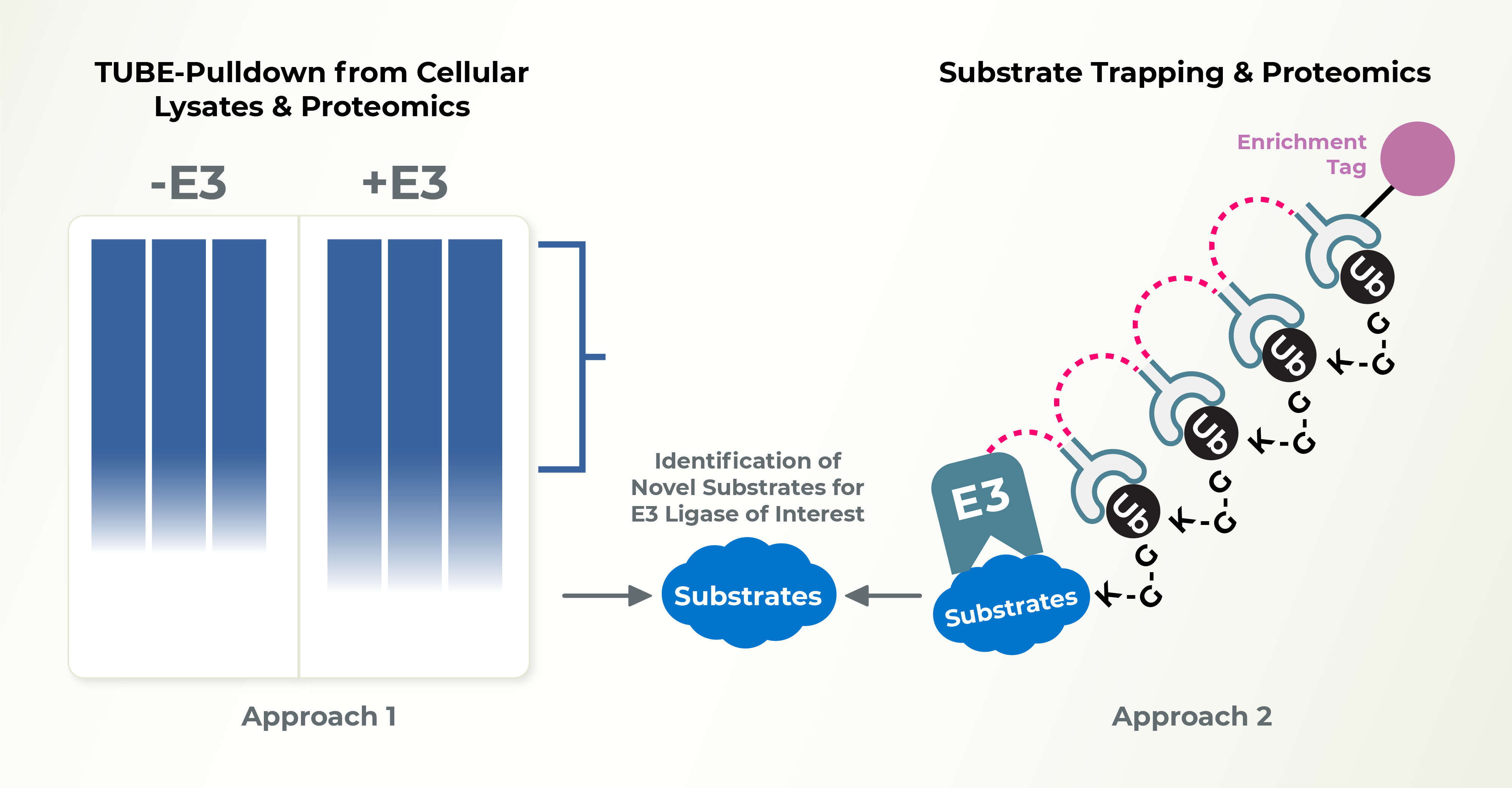
E3 Ligase Substrate ID
Recent development of PROTAC drugs to recruit E3 ligases and degrade therapeutic proteins highlight the value of studying E3 ligases. Identification of substrates for individual E3 ligases is an essential step to unravel the cellular functions of E3s and identify novel therapeutic targets. Lifesensors can help you identify the substrates of your preferred E3s.
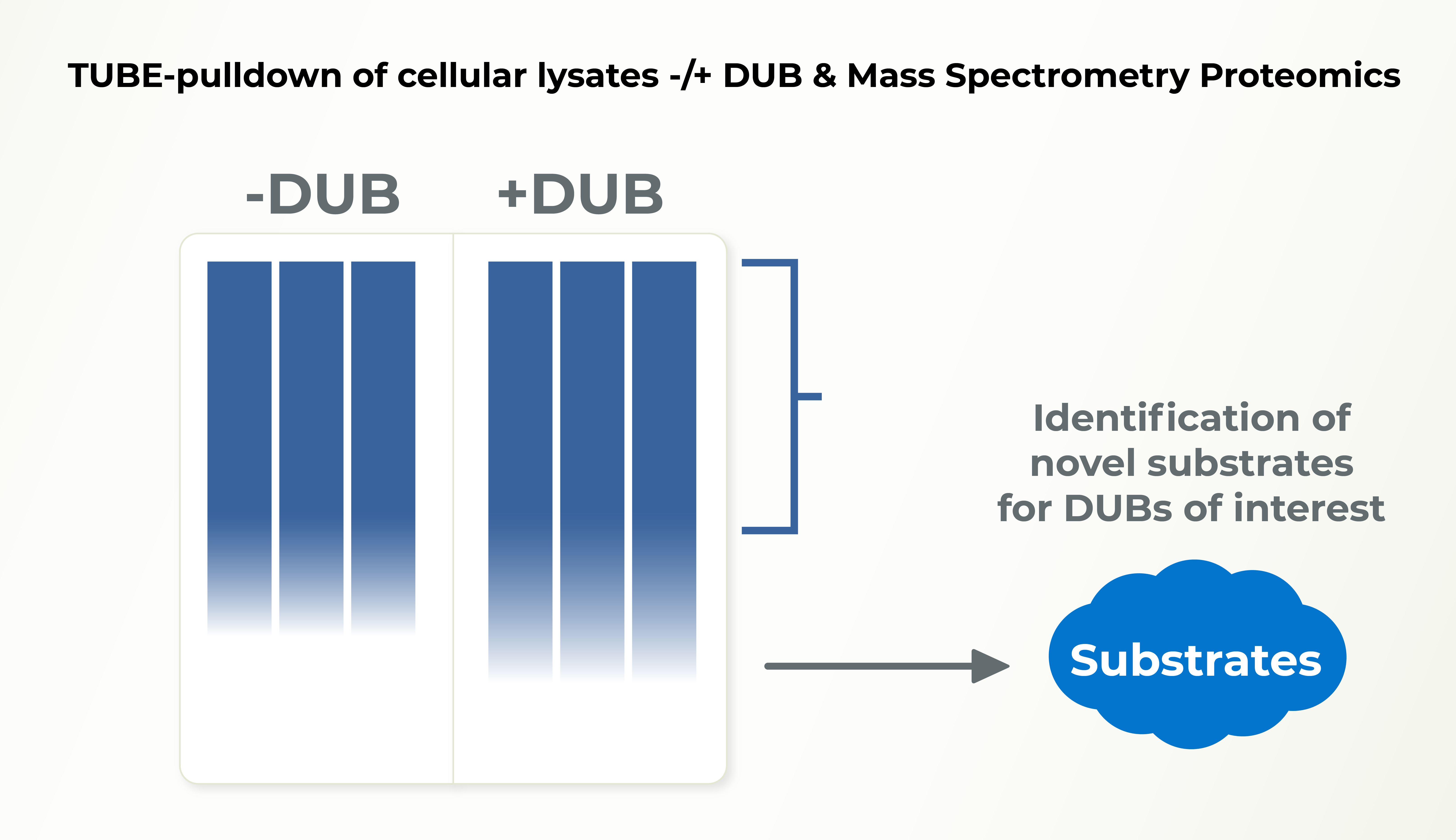
DUB Substrate ID
There are ~100 Deubiquitinases (DUBs) encoded in human genome and the functions of most of these DUBs remain unknown. Identification of substrates for individual DUBs is an essential step to unraveling their cellular functions. Lifesensors can help you identify the substrates of your preferred DUBs.
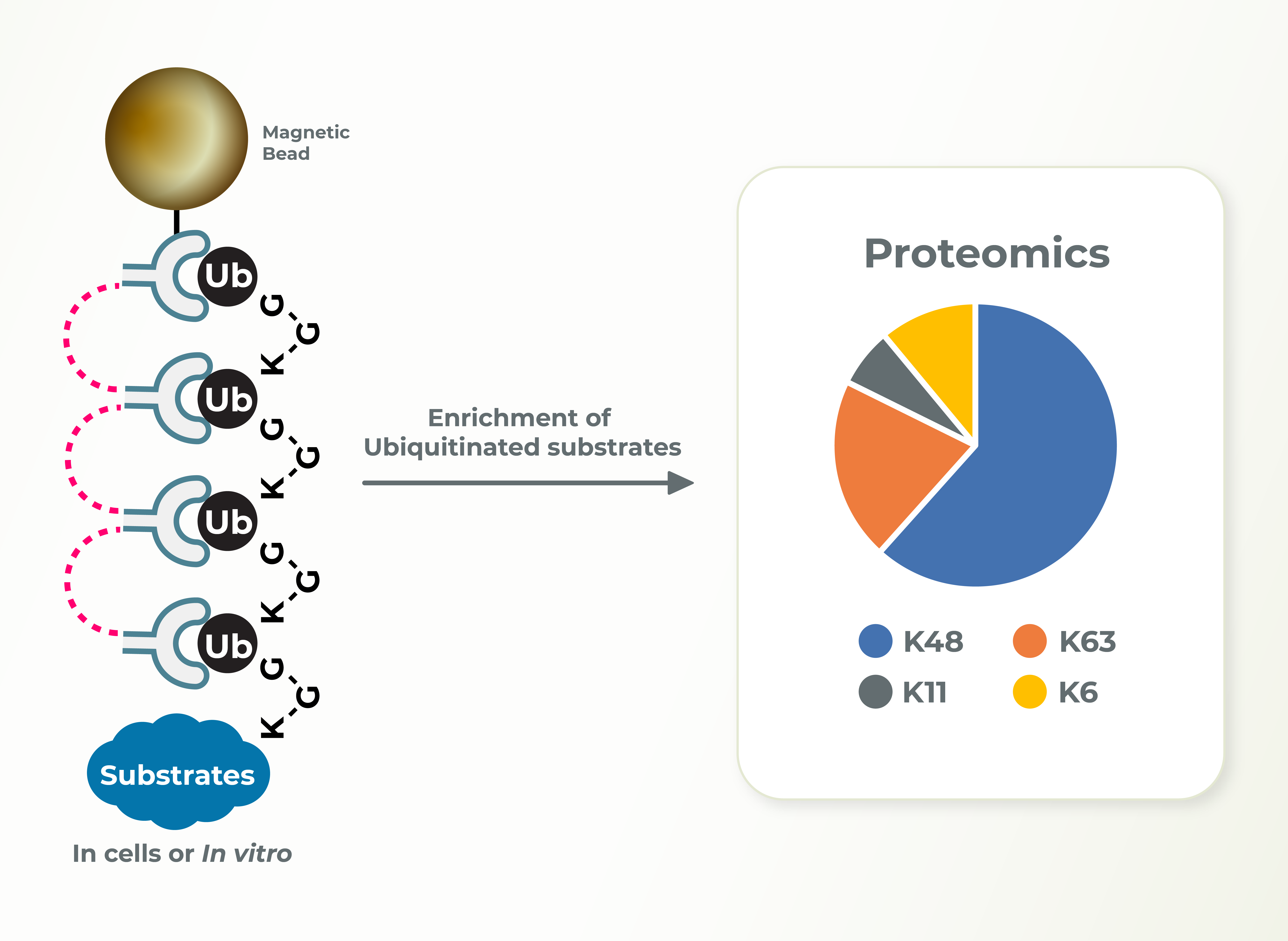
Target Substrate Lysine Ubiquitination ID
Identification of the type of ubiquitin linkage as well as the substrate lysines modified with ubiquitin can provide insights into the cellular functions of E3s. Lifesensors can perform these studies very efficiently and in a timely manner for the clients.
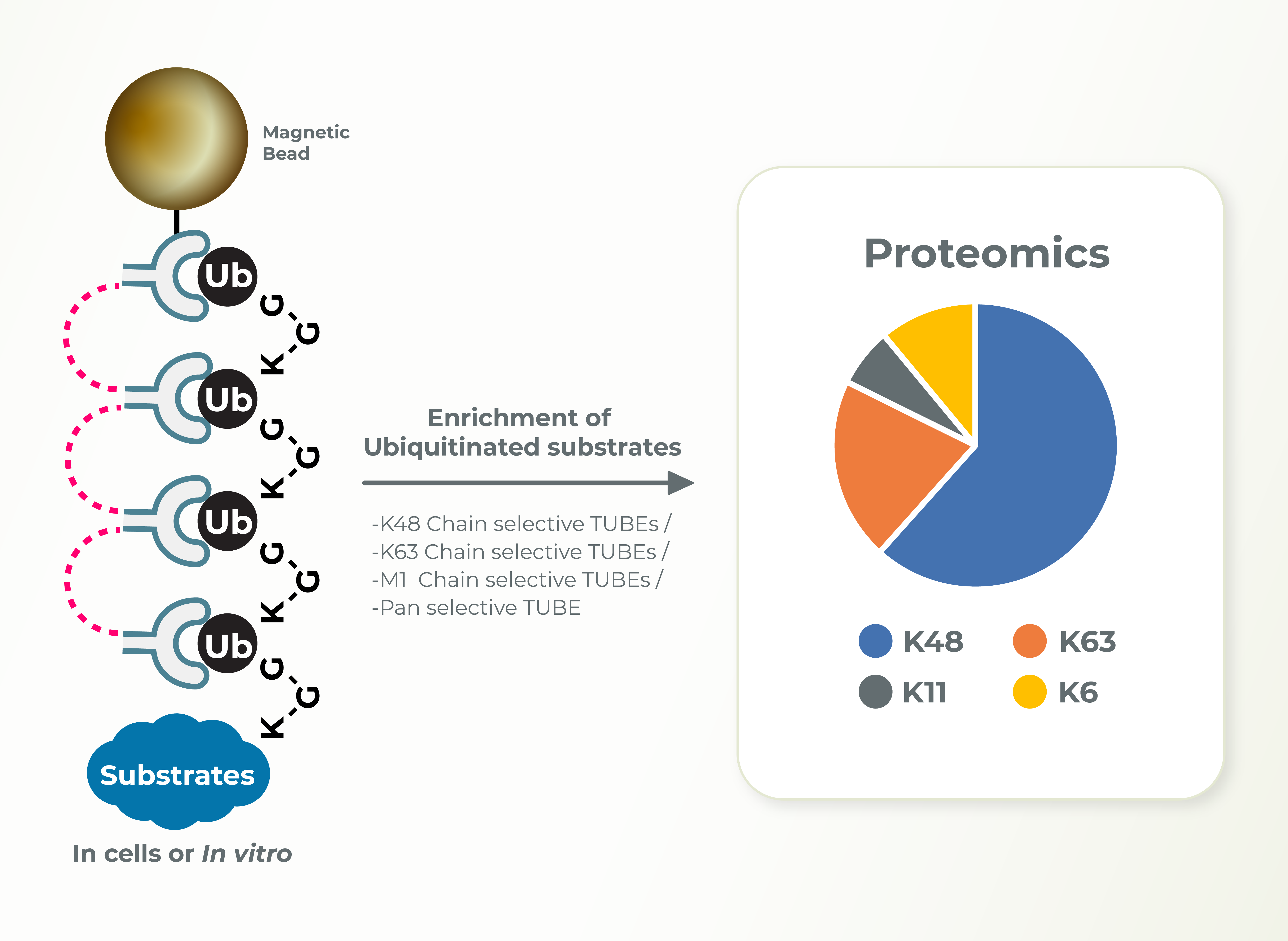
Ubiquitin Chain Diversity ID
K48-linked ubiquitin chains promote proteasomal degradation of substrates, K63-linked ubiquitin chains are associated with signaling and localization. Identification regarding the type of ubiquitin linkage can provide insights into the cellular functions of E3s and substrates. Lifesensors can perform these studies using linkage specific TUBEs very efficiently.